Complex Carbohydrate Magnetic Resonance Database (CCMRD)
Carbohydrates are crucial for various life activities in living organisms. The Complex Carbohydrates Magnetic Resonance Database (CCMRD), a solid-state NMR database for complex carbohydrates developed at Michigan State University. The Solid-state NMR spectroscopy reveals the molecular structure and dynamics of insoluble complex carbohydrates.
| Database ID | Trival Name | Linear Code | Compound Class | Taxonomy Domain | Species | Residue | SNFG | Chemical Structure | Chemical Shifts (ppm) | Spectrometer (MHz) | Temperature (K) | pH | NMR Reference Compound | Sample Treatment | Reference |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ccmrd_658 | Xyloglucan | Xylp | Polysaccharide | plant | Spruce (Picea abies) | ?-?-Xylp |
|
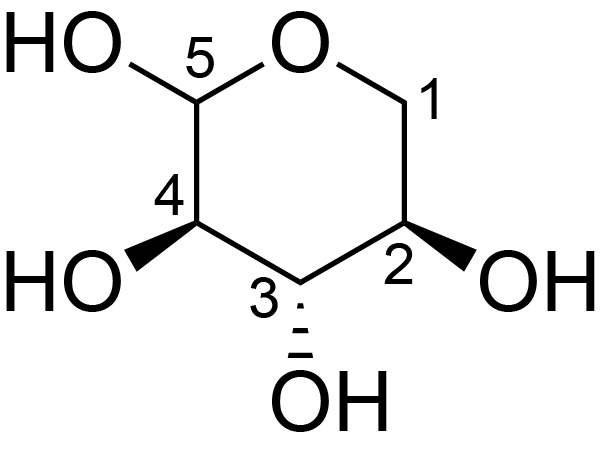
|
C1: 100.3, C2: 72.3, C4: 71.9, C5: 61.3 | 600/400 | 298.0 | None | TMS | whole cell | Alex Kirui et al., Carbohydrate-aromatic interface and molecular architecture of lignocellulose, NATURE COMMUNICATIONS, 2022 |
| ccmrd_659 | cellulose | Glcp | Polysaccharide | plant | Spruce (Picea abies) | b-D-Glcp |
|
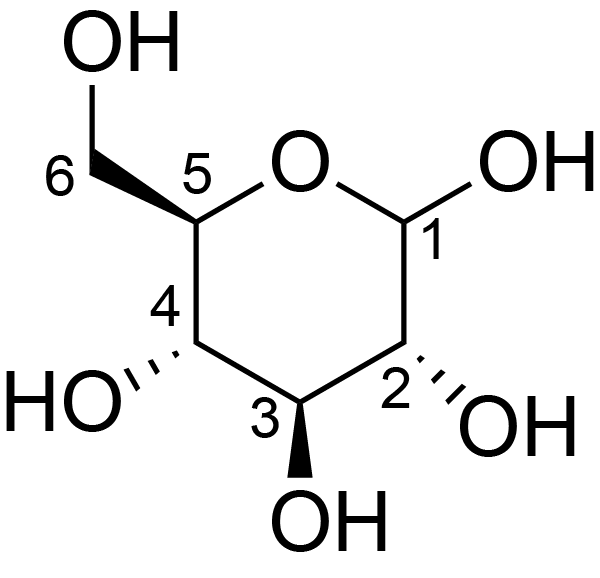
|
600/400 | 298.0 | None | TMS | whole cell | Alex Kirui et al., Carbohydrate-aromatic interface and molecular architecture of lignocellulose, NATURE COMMUNICATIONS, 2022 | |
| ccmrd_660 | Xylan | Xylp | Polysaccharide | plant | Spruce (Picea abies) | ?-?-Xylp |
|
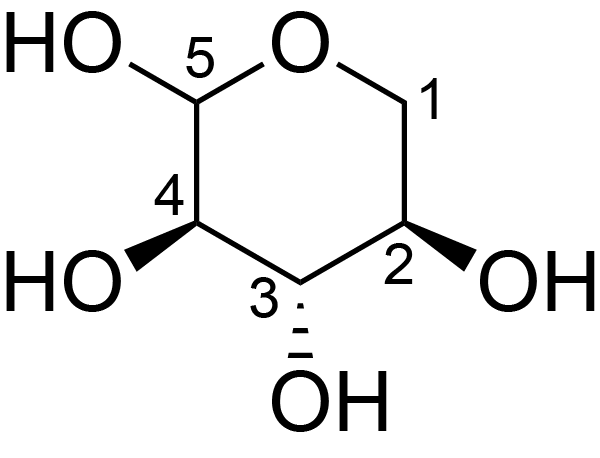
|
C1: 104.8, C2: 72.2, C3: 73.3, C4: 82.2, C5: 63.9 | 600/400 | 298.0 | None | TMS | whole cell | Alex Kirui et al., Carbohydrate-aromatic interface and molecular architecture of lignocellulose, NATURE COMMUNICATIONS, 2022 |
| ccmrd_661 | Xylan | Xylp | Polysaccharide | plant | Spruce (Picea abies) | ?-?-Xylp |
|
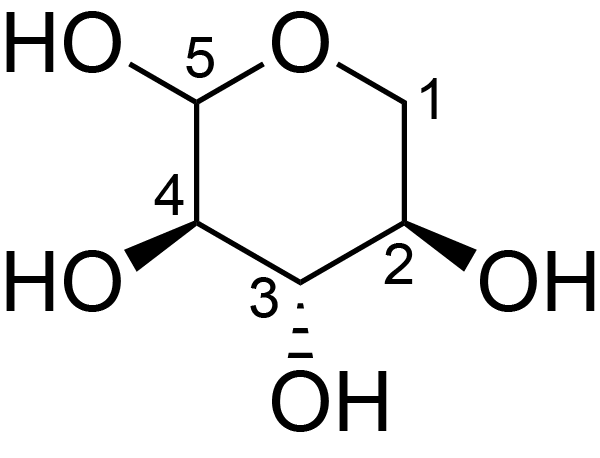
|
C1: 103.5, C2: 73.9, C3: 74.2, C4: 77.8, C5: 65.4 | 600/400 | 298.0 | None | TMS | whole cell | Alex Kirui et al., Carbohydrate-aromatic interface and molecular architecture of lignocellulose, NATURE COMMUNICATIONS, 2022 |
| ccmrd_662 | Xyloglucan | Xylp | Polysaccharide | plant | Spruce (Picea abies) | ?-?-Xylp |
|
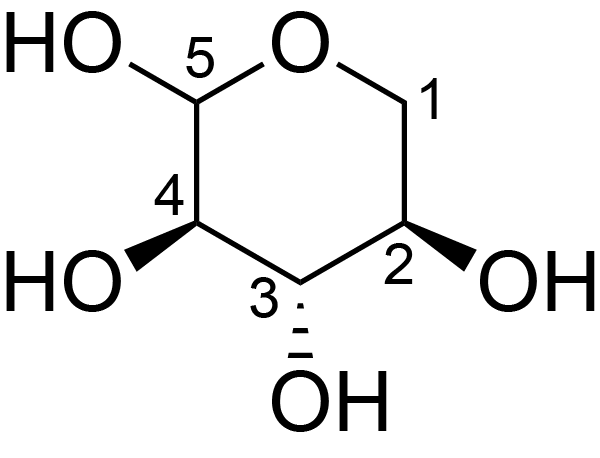
|
C3: 74.1, C4: 70.4, C5: 62.5 | 600/400 | 298.0 | None | TMS | whole cell | Alex Kirui et al., Carbohydrate-aromatic interface and molecular architecture of lignocellulose, NATURE COMMUNICATIONS, 2022 |
| ccmrd_663 | Glucuronic acid | GlcpAc | Polysaccharide | plant | Spruce (Picea abies) | ?-?-GlcpAc |
|

|
C1: 101.5, C2: 71.3, C3: 79.8, C4: 69.9, C5: 76.6 | 600/400 | 298.0 | None | TMS | whole cell | Alex Kirui et al., Carbohydrate-aromatic interface and molecular architecture of lignocellulose, NATURE COMMUNICATIONS, 2022 |
| ccmrd_664 | Glucuronic acid | GlcpAc | Polysaccharide | plant | Spruce (Picea abies) | ?-?-GlcpAc |
|

|
C1: 99.2, C2: 69.9 | 600/400 | 298.0 | None | TMS | whole cell | Alex Kirui et al., Carbohydrate-aromatic interface and molecular architecture of lignocellulose, NATURE COMMUNICATIONS, 2022 |
| ccmrd_665 | Glucuronic acid | GlcpAc | Polysaccharide | plant | Spruce (Picea abies) | ?-?-GlcpAc |
|

|
C1: 98.5, C2: 71.6, C3: 80.8 | 600/400 | 298.0 | None | TMS | whole cell | Alex Kirui et al., Carbohydrate-aromatic interface and molecular architecture of lignocellulose, NATURE COMMUNICATIONS, 2022 |
| ccmrd_666 | Galacturonic acid | GalpA | Polysaccharide | plant | Spruce (Picea abies) | ?-?-GalpA |
|
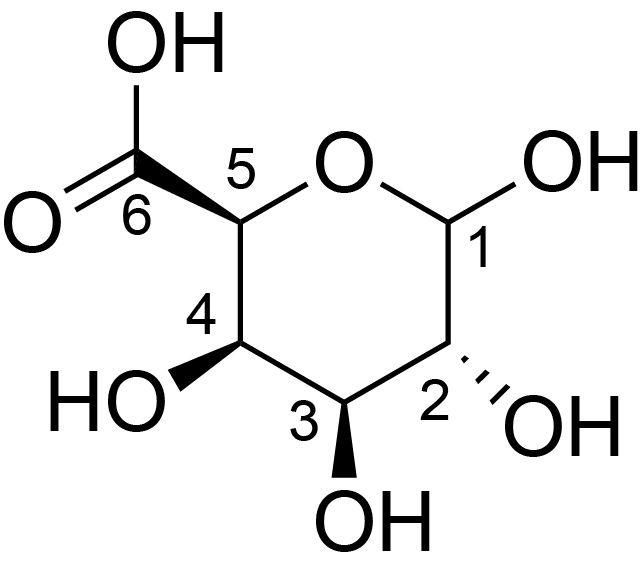
|
C1: 100.2, C2: 68.1, C4: 79.6 | 600/400 | 298.0 | None | TMS | whole cell | Alex Kirui et al., Carbohydrate-aromatic interface and molecular architecture of lignocellulose, NATURE COMMUNICATIONS, 2022 |
| ccmrd_667 | Mannan | Manp | Polysaccharide | plant | Spruce (Picea abies) | ?-?-Manp |
|
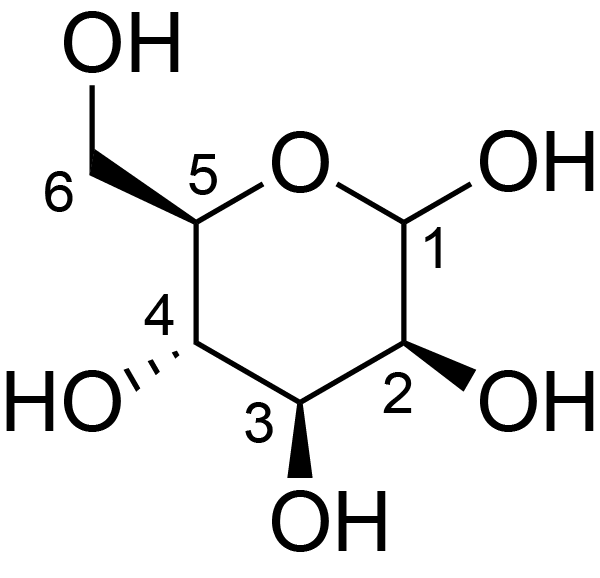
|
C1: 101.2, C2: 71.3, C3: 75.1, C5: 75.1, C6: 62.1 | 600/400 | 298.0 | None | TMS | whole cell | Alex Kirui et al., Carbohydrate-aromatic interface and molecular architecture of lignocellulose, NATURE COMMUNICATIONS, 2022 |